C5.1: Modeling Assembly and Growth of Materials Based on DNA-Hybrids
Subproject Leader: Wolfgang Wenzel
Institute of Nanotechnology (INT), KIT
Contributing Scientists:
Present: Irene Meliciani
Past: Konstantin Klenin, Horacio Perez-Sanchez, Aina Quintilla
Towards tailored materials
In most naturally occurring materials the arrangement of the constituents, be it atoms or molecules, is controlled to the rules of chemistry, which go back to the quantum mechanical principles that were discovered in the last century. Novel fabrication methods now permit the design of a bewildering array of novel nanoscale building blocks that interact according to different principles. The goal of this project is the design of materials with building blocks that interact with each other by exploitation of the pairing principles of DNA that also organize their genetic information. DNA molecules consist of subunits labeled by the letters A, T, C and G which can be synthesized in any particular sequence. Because the subunits A and T, as well as C and G, can bind to each other, respectively, DNA molecules with matching complimentary sequences can form complexes that selectively link just the matching molecules. By designing sets of complementary DNA molecules that are connected to each other one can design specific bonding patterns that result in predefined geometric assemblies. While this has been realized in a number of spectacular assemblies of finite size, the design of extended materials, e.g. crystals, based on this principle is still in its infancy. Realization of such materials would permit control of material properties at the nanoscale, such as spacing and coordination, with many exciting applications. Because the DNA is ready fabricated and such materials self-assemble realization of this goal would put a wide range of novel materials within the reach.
Simulation challenges
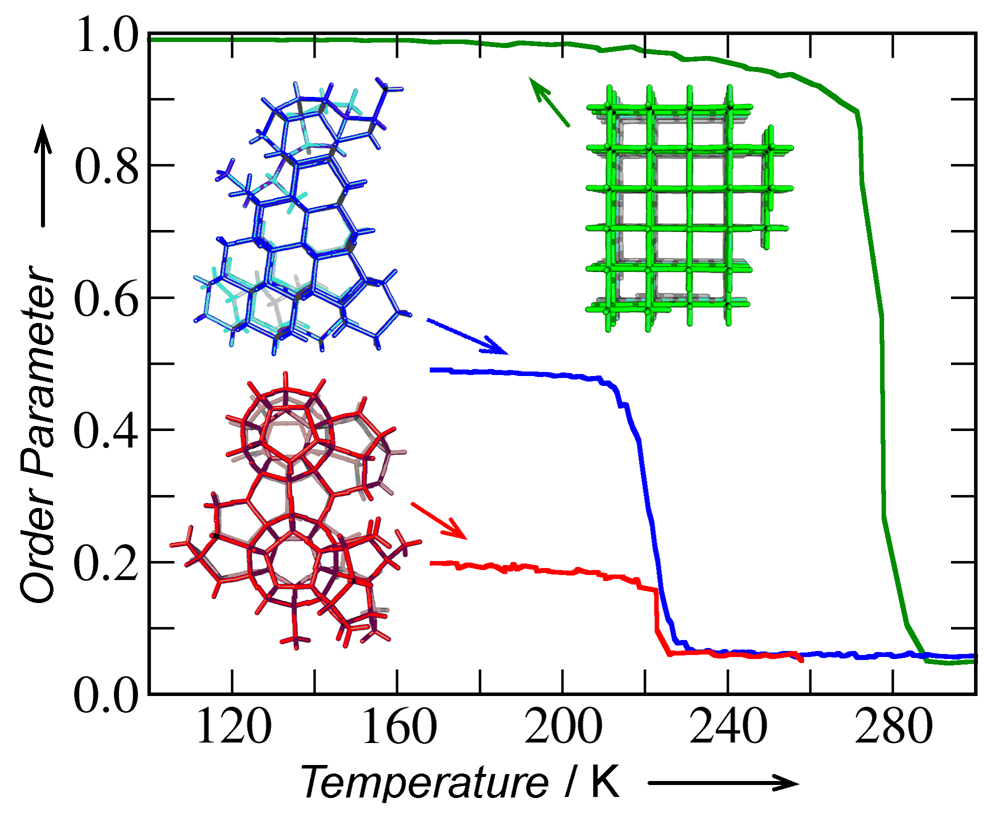
Not surprisingly a great number of factors influence the growth and assembly processes of such novel materials. It is not known a priori whether and how the building blocks fit together in space, how the interactions that are well understood for individual DNA molecules, change in the presence of so many other molecules and how faults and defects that invariably arise in the growth of any material can be subsequently heal into defect-free materials. Because novel interactions are exploited, little it is presently known about the behavior of such materials at the nano- and mesoscale and their growth processes. Simulations may therefore help to guide and accelerate the design process, avoid costly experiments and help us understand the rules of how such materials can be made. Unfortunately the number of atoms in even small fragments of such materials is so large that it is impossible to simulate the growth process using atomistic methods. We have therefore developed a multiscale approach that extracts key parameters from atomistic simulations of the constituents of such materials and integrates[1, 2] them into coarse-grained simulations for the assembly of novel materials. Using these methods we could understand how the coordination and flexibility of the DNA sequences influences the assembly properties of the resulting materials and support our experimental colleagues in the design of novel DNA-based materials that are stable almost to the boiling point of water.
Supporting other CFN efforts
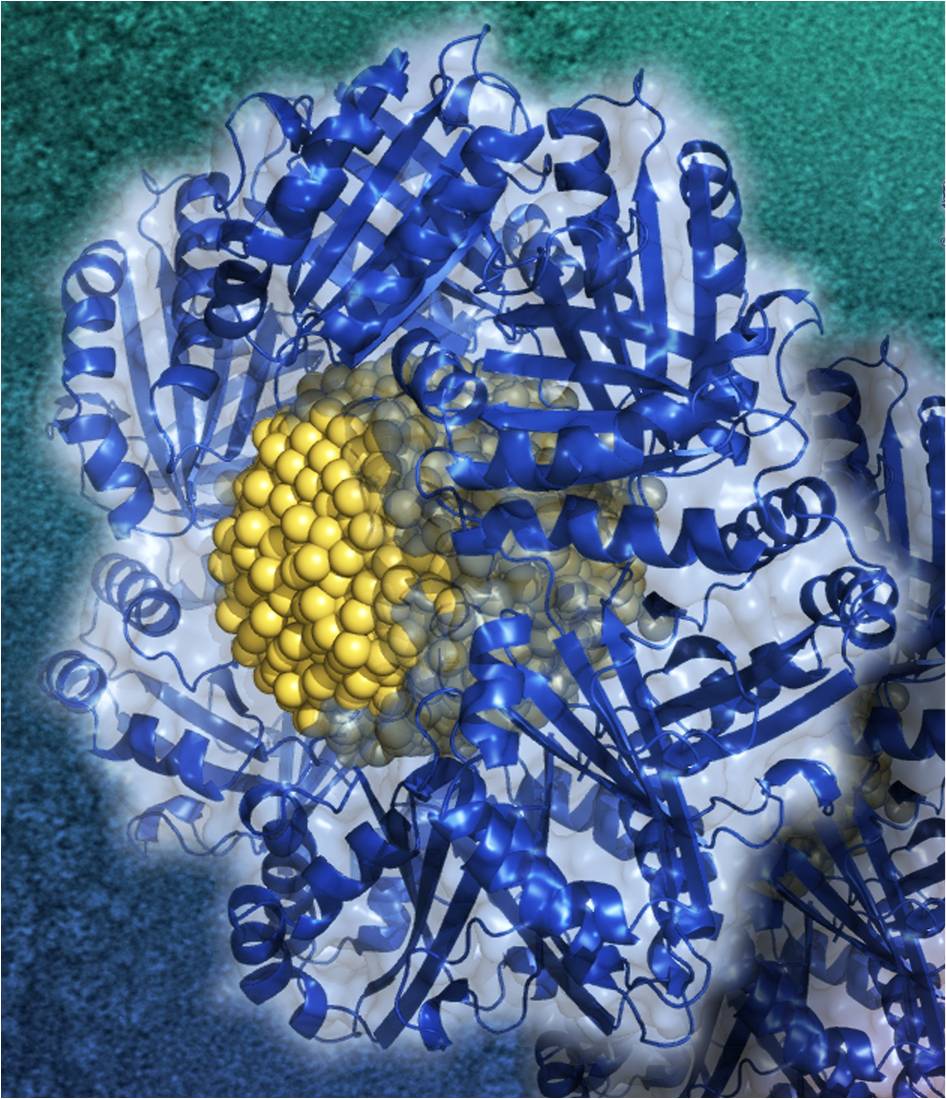
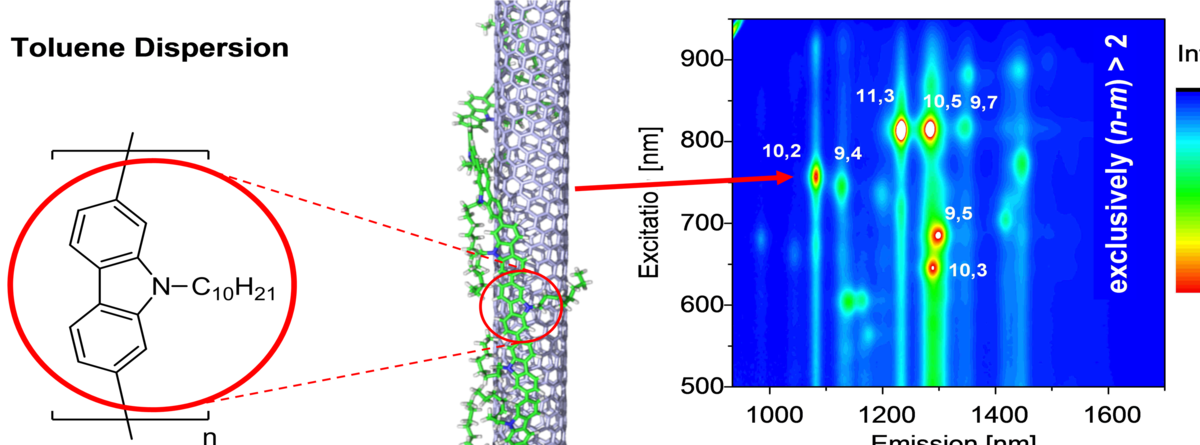
Using these methods we could also contribute to the elucidation of other fascinating nanoscale growth processes, such as realization of when electrochemically gated atomic switch will with predefined quantum conductance[3], the controlled growth of catalytically active metallic nanoparticles into protein templates[4] that self-assemble into dendritic structures and the development of novel techniques to sort carbon nanotubes by size using specific polymers that selectively bind to a particular species of carbon nanotube[5].
|
[1] |
Meng, M., et al., Two Base Pair Duplexes Suffice to Build a Novel Material. ChemBioChem, 2009. 10(8): p. 1335-1339. |
|
[2] |
Singh, A., et al., Branched DNA That Forms a Solid at 95 degrees C. Angewandte Chemie-International Edition, 2011. 50(14): p. 3227-3231. |
|
[3] |
Xie, F.Q., et al., Independently Switchable Atomic Quantum Transistors by Reversible Contact Reconstruction. Nano Letters, 2008. 8(12): p. 4493-4497. |
|
[4] |
Behrens, S., et al., Constrained Synthesis and Organization of Catalytically Active Metal Nanoparticles by Self-Assembled Protein Templates. Advanced Materials, 2009. 21(34): p. 3515. |
|
[5] |
Lemasson, F.A., et al., Selective Dispersion of Single-Walled Carbon Nanotubes with Specific Chiral Indices by Poly(N-decyl-2,7-carbazole). J Am Chem Soc, 2011. 133(4): p. 652-655. |
List of Publications 2006-2011 as PDF
Subproject Report 2006-2010 as PDF